Sponsored by the Cushing/Whitney Medical Library
Date & Time: 9:00am - 12:30pm, Friday, June 5, 2015
Location: The Anlyan Center Auditorium (N 107), 300 Cedar Street New Haven, CT 06520
Campus: Medical School
Presenter: Dr. Alex Kaplun, Field Applications Scientist, BIOBASE
Registration: Free and open to Yale affiliates – limited seating-
PROTEOME™’s powerful ontology search query system, with specialized tools for gene set analysis and pathway visualization, allows scientists to quickly find answers to questions relevant to their research. It works seamlessly with TRANSFAC®, an internationally unique knowledgebase containing data on eukaryotic transcription factors and miRNAs, their experimentally-proven binding sites, and regulated genes, which supports research into gene regulation. Based on TRANSFAC®'s broad compilation of binding sites, positional weight matrices are derived which can be used with the included Match tool to search DNA sequences for predicted transcription factor binding sites. TRANSFAC enables you to identify transcription factors affecting gene expression in your microarray and RNA-Seq experiments, as well as predict how they, in combination, can induce observed gene expression patterns.
In the PROTEOME™ section, the attendees will learn to:
1. Search for individual gene, disease, and drug reports by name.
2. Browse for sets of genes, diseases, and drugs which share a desired set of characteristics.
3. Upload a list of genes and identify those characteristics which are statistically over-represented (NEW)
4. Export annotated characteristics for a gene list.
5. Visualize protein-protein networks, overlaid with disease and drug assignments
6. Annotate custom sequences.
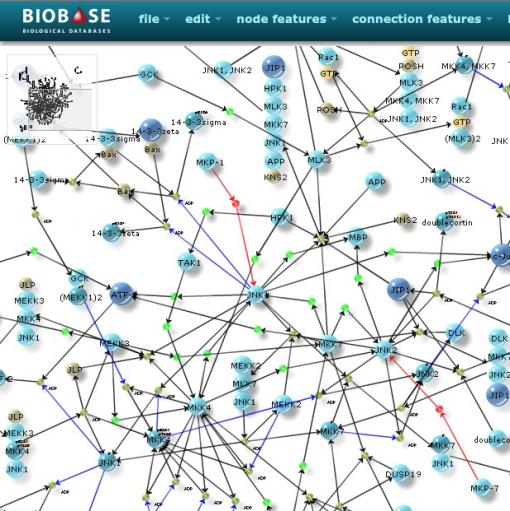
Network visualization using the BKL Pathfinder tool.
In the TRANSFAC section, the attendees will learn to:
1. Search for individual transcription factors and miRNAs, their experimentally-characterized binding sites and regulated genes, and ChIP experiments.
2. Create positional weight matrices of transcription factor binding sites using set of aligned experiment-derived sites.
3. Predict transcription factor binding sites (single sites or combinations) within a promoter or DNA sequence.
4. Analyze high-throughput data sets for models of transcription factor binding (NEW).
5. Perform statistical analysis of your differential expression data to determine which transcription factors are responsible for the observed effect (NEW).
6. Perform step-by-step comprehensive microarray and ChIP-seq data analysis in easy-to-use, guided workflows (NEW).