The National Center for Biotechnology Information is developing a new type of BLAST called SmartBLAST. It process the user query in such a way that presents the three best matches from the non-redundant protein sequence database along with the two best protein matches from well-studied reference species. In addition, it provides results that match the query from the Conserved Domain Database (CDD)
SmartBLAST accepts only one query at a time- either as FASTA sequence or protein accession number/GI- and uses a combination of BLAST and a multiple sequence alignment to produce its results. It first uses the query to search the non-redundant (nr) protein database. Then, it searches the reference database with BLASTP, followed by a multiple sequence alignment on the six sequences (the query and five subject sequences) using the COBALT multiple sequence alignment program.
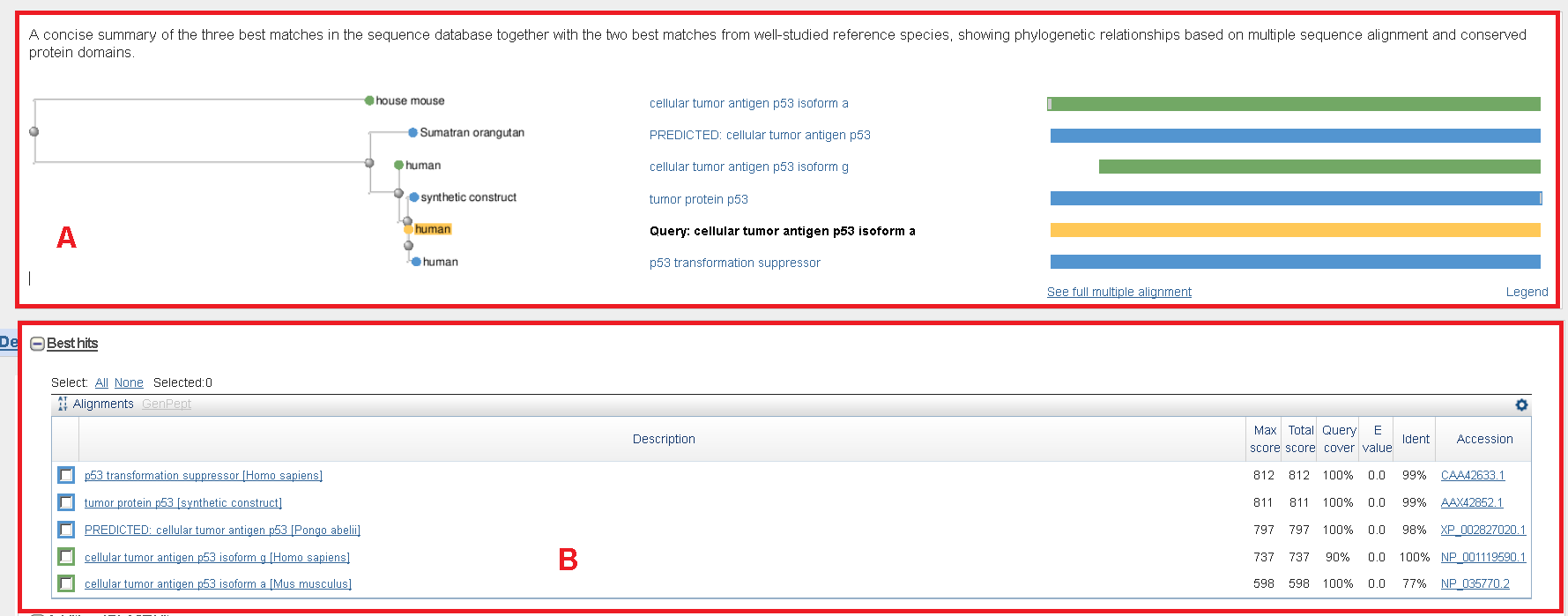
Screen capture showing the results of a SmartBLAST for TP53 (GI:187830777). Panel A shows the five matching sequences are represented as a phylogenetic tree and a graphical overview. The matches are color-coded: matches from the reference species are green, matches from the non-redundant protein database are blue, and your query is yellow. Panel B represents the results from the multiple alignments.
Join the End-User Bioinformatics Network (EBNET) and become a member of a grass root community that collaborates on end-user bioinformatics events, training sessions, resources, and tools that support biomedical research at Yale.