The End-user Bioinformatics Program at the Yale Cushing/Medical Library is hosting these four workshops on tools for the analysis of NGS data. Besides the two trainings on tools for functional analysis of NGS data already supported by the Medical Library (Ingenuity Pathway Analysis and MetaCore), we will have a presentation on Partek Flow, and another one on CLC Bio (QIAGEN). The medical library will ask for feedback on these tools for future support and licensing. If you are interested in attending any of these presentations, please register to reserve your seat. Please contact Rolando Milian for questions or comments.
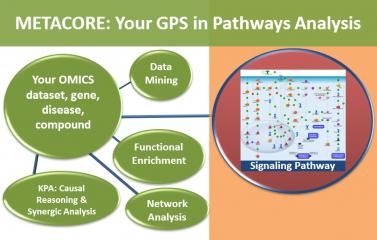
Title: Introductory Workshop to MetaCore and Key Pathway Advisor – Pathway Analysis of “Omics” Data
This hands-on training workshop will highlight basic functionalities as well as cover use cases to:
- Predict upstream regulators of gene expression using causal reasoning
- Use synergic enrichment analysis of upstream regulators and observed gene expression changes to identify key pathways associated with your data
- Compare between experiments to uncover phenotypic differences using enrichment analyzes
- Search and explore genes, proteins, diseases and compounds
Date & Time: 9:00am - 11:00am, Thursday, April 28, 2016
Location: Beaumont room (2nd floor), SHM, 333 Cedar St, New Haven CT 06510
Presenter: Deborah Riley, PhD, Senior Solution Scientist – Thomson Reuters Life Sciences
Title: Start-to-finish Analysis Software for NGS & Microarray Data.
(miRNA-Seq Analysis with Partek: Serum miRNA Study in Alcohol Use Disorder Subjects Suggests Alterations of CNS Structure and Function)
During this seminar, we will feature a successful miRNA-Seq based study of extracellular miRNAs in 20 individuals diagnosed with alcohol use disorder (AUD). We will demonstrate how to go from raw NGS data to biological interpretation using Partek software. Analysis of the sequencing data using Partek Flow will include:
- checking the quality of reads
- generating aligned reads
- quantifying miRNA levels
- determining differentially expressed miRNAs
By integrating miRNA-Seq results in Partek Genomics Suite, we will demonstrate how differentially expressed miRNAs impact CNS structure and function using Partek Pathway. Lastly, miRNA expression microarray data will be analyzed in Partek Genomics Suite to validate findings from the next generation sequencing data.
Date & Time: 9:00am - 11:00am, Tuesday, May 3, 2016
Location: C-103 - SHM 333 Cedar St, New Haven CT 06520
Presenter: Dr. Eric Seiser, Field Application Scientist, Partek Incorporated

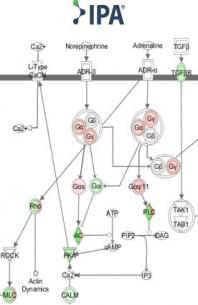
Title: Ingenuity Pathway Analysis Hands On Training
If you have gene (including RNAseq), protein and metabolic expression data, you should be using IPA to guide you with the biological interpretation of your data. Using IPA you will learn how to rapidly understand:
- Pathway involvement and change
- Effected biological processes
- Causal regulators and their directional effect on genes, functions and diseases across multiple time points or doses. You will also learn to explore IPA’s knowledge and discovery tools that allow you to relate the most recent literature findings to your research.
Requirement: Laptop and active IPA account (Request your account here)
Presenter: Devendra Mistry, PhD, Field Application Scientist, Ingenuity Products, QIAGEN
Date & Time: 9:00am - 11:30am, Tuesday, May 10, 2016
Location: C-103 - SHM 333 Cedar St, New Haven CT 06510
Title: CLC Genomics Workbench
Overview of Application, Importing NGS read data, QC & Pre-processing
- De novo assembly – Genomes & Transcriptomes. Characterizing Contigs, Joining & Finishing
- Mapping/Alignment to Reference, Variant Calling, Annotation & Filtering
- RNA Seq Analysis Workflow & Tools
- Overview of Microbial Modules (Finishing & Microbial Genomics)
- CLC Biomedical Workbench & Ingenuity Variant Analysis
- Prebuilt intuitive pipeline for your human DNA-seq data that allows you to quickly go from reads or called variants to identifying and prioritizing the casual variants.
Date & Time: 1:00pm - 3:00pm, Tuesday, May 10, 2016
Location: C-103 - SHM 333 Cedar St, New Haven CT 06510
Presenter: Devendra Mistry, PhD, Field Application Scientist, Ingenuity Products, QIAGEN