On April 5 and 6, Dr. Peter Cooper*** will provide training in the form of four workshops (see below) on the some of the most valuable National Center for Biotechnology Information bioinformatics resources and tools at Yale School of Medicine. This training is hosted by the Yale Cushing/Whitney Medical Library. Although free and open to any Yale affiliate, it is recommended to register since seating is limited.
Please contact Rolando Milian for questions on these sessions: 203-785-6194
A Practical Guide to NCBI BLAST
This workshop highlights important features and demonstrates the practical aspects of using the NCBI BLAST service, the most popular sequence similarity service in the world. You will learn about useful but under-used features of the service. These include access from the Entrez sequence databases; the new genome BLAST service quick finder; the integration and expansion of Align-2-Sequences; organism limits and other filters; re-organized databases; formatting options and downloading options; and TreeView displays. You will also learn how to use other important sequence analysis services associated with BLAST including Primer BLAST, an oligonucleotide primer designer and specificity checker; the multiple protein sequence alignment tool, COBALT; IgBLAST, a tool for analysis of antibody and T-cell receptor sequences; and MOLE-BLAST, a new tool for clustering and providing taxonomic context for targeted loci sequences (16S, ITS, 28S). These aspects of BLAST provide easier access and results that are more comprehensive and easier to interpret.
Date: Tuesday, April 5, 2016
Time: 9:00am - 12:00pm
Location: C-103 - SHM 333 Cedar St, New Haven CT 0652
Accessing Genomes, Assemblies and Annotation Products
You will learn how NCBI processes genome-level data and produces annotation through the prokaryotic and eukaryotic genome annotation pipelines. You will find, browse, and download genome-level data for your organism of interest and for environmental and organismal metagenomes using the Genome, BioProject and Assembly resources. In addition to assembled and annotated data, you will retrieve and download draft whole genome shotgun and read-level next-gen sequencing data from the Nucleotide and Sequence Read Archive (SRA) databases. You will access results of precomputed analyses of genomes, as well as perform your own analyses of assembled and unassembled genomic data using NCBI's genome BLAST and SRA-BLAST services.
Date: Tuesday, April 5, 2016
Time: 1:30pm - 4:00pm
Location: C-103 - SHM 333 Cedar St, New Haven CT 06520
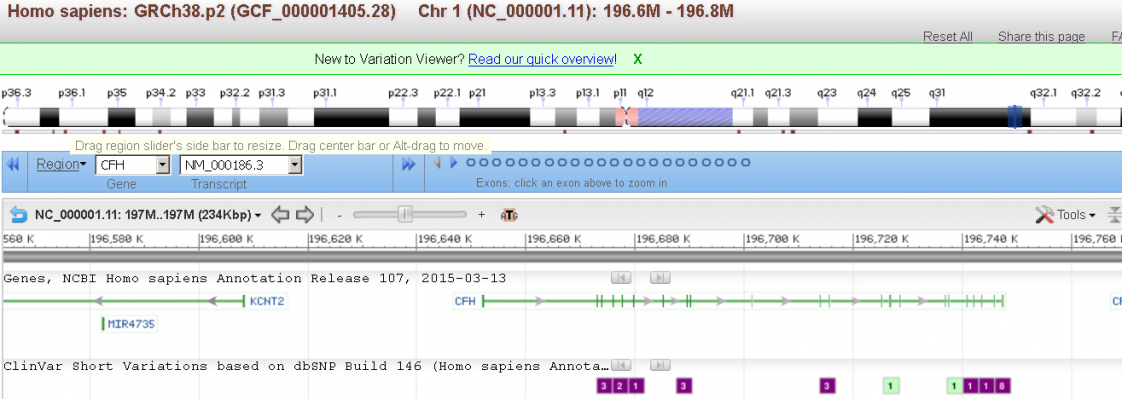
Accessing NCBI Human Variation and Medical Genetics Resources
You will learn to use and access resources associated with human sequence variations and phenotypes associated with specific human genes and phenotypes. The workshop will emphasize the Gene, MedGen and ClinVar resources to search by gene, phenotype and and variant respectively. You will learn how to map variation from dbSNP and dbVAR onto genes, transcripts, proteins, and genomic regions and how to find genetic tests in GTR. You will also gain experience using additional tools and viewers including PheGenI, a browser for genotype associations and the new Variation Viewer the 1000 Genomes Browser, which provide a useful ways to search for, map and browse variants as well as upload and download data in genomic context.
Date: Wednesday, April 6, 2016
Time: 9:00am - 12:00pm
Location: C-103 - SHM 333 Cedar St, New Haven CT 06520
Exploring Gene Expression Information at the NCBI
You will find, display and analyze microarray and sequence-based expression data that are stored in the Gene Expression Omnibus (GEO), Sequence Read Archive (SRA), UniGene, and Epigenomics databases to investigate the potential for expression of transcript splice variants and examine the levels of expression under varied experimental conditions as well as in different tissues and disease states. You will analyze Microarray data the on-demand GEO2R tool and will explore the precomputed transcript analyses that are displayed on the UniGene and GEO Profiles pages. You will explore genome-aligned RNA-Seq data through the Gene database's sequence viewer displays and analyze raw RNA-Seq reads in the SRA database using NCBI's SRA-BLAST service.
Date: Wednesday, April 6, 2016
Time: 1:30pm - 4:00pm
Location: C-103 - SHM 333 Cedar St, New Haven CT 06520
***Dr. Peter Cooper, Staff Scientist, National Center for Biotechnology Information (NCBI) directs the scientific outreach and training program for the National Center for Biotechnology Information at the National Library of Medicine. Peter has conducted and developed training courses for biologists in the use of NBCI molecular databases and has provided scientific user support for the NCBI since 1998. Prior to joining the NCBI Peter pursued diverse biological research interests including peptide neurochemistry, marine environmental toxicology, and taught biology and chemistry. Peter earned a BS from Virginia Tech, a MA in chemistry from the Johns Hopkins University and a Ph.D. in Marine Science from the College of William and Mary, School of Marine Science in 1996